Metabolomics in sepsis-associated acute kidney injury research
D.Ivšić Škoda, D. Rešetar Maslov*, J. Kuleš, V. Benko, V. Đurić, R. Barić Rafaj and V. Mrljak
Dora Ivšić Škoda1, divsicskoda@vef.unizg.hr; Dina Rešetar Maslov1* (corresponding author), drmaslov@vef.unizg.hr, orcid.org/0000-0002-0159-8281; Josipa Kuleš2, jkules@vef.unizg.hr, orcid.org/0000-0002-4945-4690; Valerija Benko3, vbenko@vef.unizg.hr, orcid.org/0000-0002-3307-9582; Vlasta Đurić1 vduric@vef.unizg.hr; Renata Barić Rafaj2, rrafaj@vef.unizg.hr, orcid.org/0000-0002-1043-1176; Vladimir Mrljak1, vmrljak@vef.unizg.hr, orcid.org/0000-0002-0723-7815.
1 Internal Diseases Clinic, Faculty of Veterinary Medicine, University of Zagreb, 10000 Zagreb, Croatia
2Department of Chemistry and Biochemistry, Faculty of Veterinary Medicine, University of Zagreb, 10000 Zagreb, Croatia
3Department for Biology and Pathology of Fish and Bees, Faculty of Veterinary Medicine, University of Zagreb, 10000 Zagreb, Croatia
https://doi.org/10.46419/cvj.57.4.3
Abstract
Metabolomics is a scientific discipline focused on the analysis of metabolites in biological samples with the aim of understanding physiological and pathological processes. The development of analytical technologies, such as mass spectrometry and nuclear magnetic resonance, has enabled advances in untargeted and targeted metabolomic analysis. In untargeted analysis, a broad spectrum of metabolites is analysed without prior assumptions, whereas targeted analysis focuses on the quantification of known molecules. Biological samples such as serum, plasma and urine are most commonly used, with the choice depending on the clinical context and the nature of the disease. Despite its potential, metabolomics faces challenges such as technical complexity, biological variability, lack of standardisation, and the need for advanced statistical methods. However, in the last decade, it has been used extensively in the study of sepsis and associated acute kidney injury, a condition with high mortality and limited therapeutic options. In this review, we present metabolomics, the challenges it faces, and the current state of research concerning sepsis-associated acute kidney injury (SA-AKI). Metabolomic studies of SA-AKI have identified potential key biological changes, including enhanced inflammatory signalling, oxidative stress, osmolyte disturbances, and cellular metabolic dysfunction. Although individual metabolites have been recognised as potential diagnostic markers, none can fully capture the complex metabolic changes characteristic of SA-AKI. In the future, the integration of metabolomics with other “omics” and advanced analytical approaches could advance early disease detection, the understanding of pathophysiology, and the development of personalised therapies.
Key words: metabolomics; metabolome; metabolites; acute kidney injury; sepsis
Metabolomics
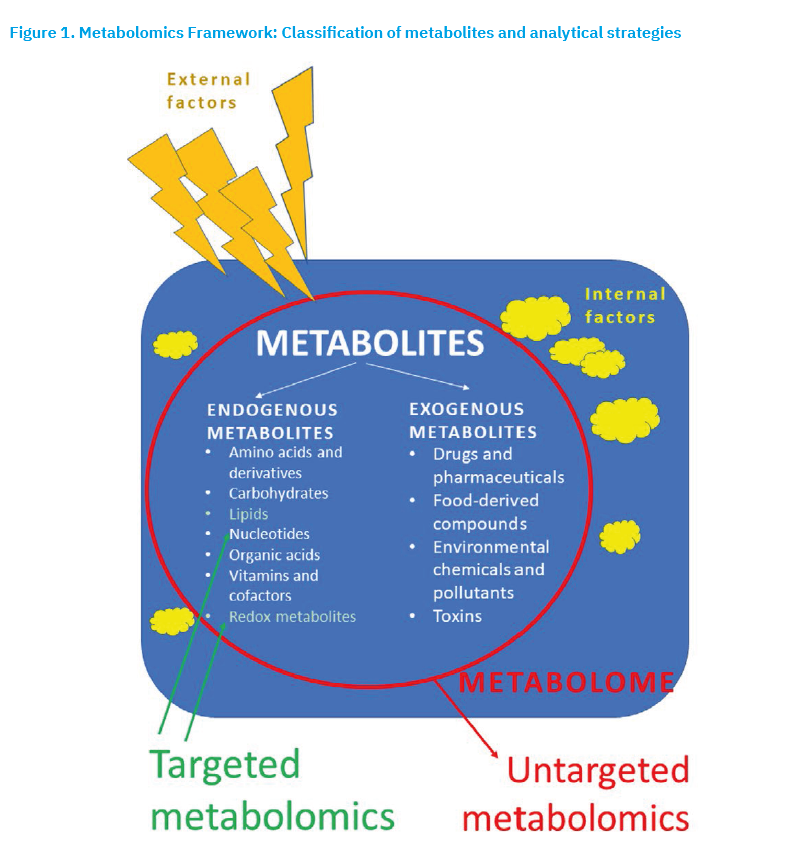
Metabolomics is a scientific discipline that allows the comprehensive analysis (detection, identification and quantification) of metabolites in a single experiment. It focuses on the metabolome, the (almost complete) set of small molecules (metabolites) present in cells, biofluids, tissues, organs or entire organisms at a given time (Hollywood et al., 2006). Metabolomics primarily analyses metabolites with a molecular mass of less than 1.5 kDa (Figure 1) (Psychogios et al., 2011; Wishart et al., 2006), but also larger metabolites exceeding 2 kDa such as complex lipids and peptides (Li-Gao et al., 2021). Endogenous metabolites are the product of cellular metabolism and the result of gene activity, while exogenous metabolites are introduced into the body from external sources (e.g., via food, drugs, toxins) (Figure 1). Endogenous metabolites are essential for normal metabolic processes, while exogenous metabolites can be either beneficial (e.g., vitamins, antioxidants) or harmful (pesticides and alcohol) (Figure 1) (Kong and Hernandez-Ferrer, 2020). Metabolomic analyses are increasingly focussing on metabolites produced by the microbiome, the community of microorganisms inhabiting specific body sites (e.g., the gut).
Metabolites reflect factors such as lifestyle and environment (diet, physical activity, medications and drugs, toxins, stress and sleep patterns), biological factors (genetics, age, gender, and circadian rhythms) and health status (pathophysiological factors like metabolic disease, inflammation, or hormonal imbalances) (Hajnajafi and Iqbal, 2025). Metabolomics is closely linked to both the phenotype (observable characteristics of the organism) and endotype (biological subtype defined by distinct metabolic pathways) of a disease (Wu et al., 2023). Because metabolites respond dynamically to internal and external factors (Figure 1), metabolomics is often considered a bridge between genotype (the totality of all genes of an organism that determine its hereditary characteristics) and phenotype, and as an important tool in personalised medicine (Jacob et al., 2019). Alterations in metabolite levels directly reflect the biochemical activity of the organism (Sindelar and Patti, 2020), providing key insights into both physiological and pathophysiological processes. Therefore, changes in metabolite levels can serve as molecular markers for early diagnosis and monitoring therapeutic responses (Vo and Trinh, 2024; Peron and Lin, 2024). Identifying specific metabolomic profiles enhances understanding of disease mechanisms, enabling the tailoring of therapies to individual patients. Like other “-omics” technologies, metabolomics is widely applied in the research of complex diseases, such as cancer, cardiovascular disease and, in particular, chronic kidney disease (Rhee, 2015). Particular emphasis is placed on conditions with high morbidity and mortality, including sepsis-associated acute kidney injury (Ping et al., 2021).
Analytical methods in metabolomic analysis
The most common analytical methods for identifying and quantifying molecules in metabolomics research are nuclear magnetic resonance (NMR) spectroscopy and mass spectrometry, often used in combination with tandem mass spectrometry (MS/MS) (Wang et al., 2010; Nagana Gowda et al., 2025). Mass spectrometry is a highly sensitive technique that is usually coupled with a separation method, such as liquid chromatography (LC) (Ovbude et al., 2024), gas chromatography (GC) (Fiehn, 2016) or capillary electrophoresis (CE) (Soga, 2023). LC-MS and GC-MS allow the analysis of a wide range of metabolites, where LC-MS is more suitable for polar and thermolabile compounds, and GC-MS is typically used for volatile metabolites. CE-MS is particularly useful for analysing highly polar and charged metabolites, such as amino acids and small peptides. NMR spectroscopy is a non-invasive analytical method that utilises the magnetic properties of selected atomic nuclei (1H, 13C, 31P) and does not require prior chromatographic separation. Other techniques, such as Fourier transform infrared spectroscopy (FTIR), Raman spectroscopy, and stable isotope-resolved metabolomics (SIRM) are used less frequently. Matrix-assisted laser desorption/ionisation time-of-flight mass spectrometry (MALDI-TOF MS) enables three-dimensional mapping of metabolites across tissue regions by analysing successive histological sections (Zhang et al., 2024).
The choice of analytical method depends on the type of sample, the physicochemical properties of the metabolites to be analysed, and the desired sensitivity. For example, high analyser sensitivity is required for detecting metabolites present at low concentrations. Unlike MS, NMR enables the quantification of metabolites without the use of standards, but is less sensitive than MS (Rubim de Santana et al., 2022). Biological samples are highly complex, encompassing a wide dynamic range of metabolite concentrations and necessitating selective methods for both sample preparation and metabolite detection. In addition to instrumental techniques, different extraction and sample preparation methods are required for polar and non-polar metabolites (Hajnajafi and Iqbal, 2025). Moreover, factors such as diet, lifestyle, environment and circadian rhythms can significantly influence the metabolomic profile, and certain metabolites are particularly susceptible to degradation. Therefore, sample collection and storage must be carefully planned and controlled (Emwas et al., 2025; de Souza et al., 2024). Metabolomics generates large volumes of complex data that require advanced statistical and bioinformatic methods for proper interpretation (Collins et al., 2021). While changes in metabolite levels can be detected, their functional significance in relation to health and disease is not always clear and often requires additional functional studies.
Metabolomic approaches
Metabolomic profiling is an analytical procedure used to measure and analyse metabolites in biological samples (Clarke and Haselden, 2008). Its aim is to characterise all, or at least most, of the detectable metabolites present in a sample. The process involves the detection, identification and quantification of metabolites, culminating in the creation of a metabolomic profile. This profile can be generated through untargeted and/or targeted metabolomic analyses (Figure 1).
An untargeted approach aims to provide a comprehensive analysis of the metabolome without prior selection of metabolites (Bingol, 2018). It is important to note that if the concentration of a metabolite falls below the method’s limit of detection (LOD), the instrument will not be able to distinguish its signal from background noise (Do et al., 2018; Beger et al., 2024). Untargeted analysis offers broad insight into the metabolomic profile and enables the discovery of new molecular markers or previously unknown metabolic pathways. It is often applied in hypothesis-free research, where the list of metabolites is not known in advance and the objective is to understand the entire metabolome. In this approach, the relative intensities of all measurable analytes, including unknown compounds, are recorded, providing valuable information for comparing of metabolome across groups (e.g., healthy vs. diseased, or between different conditions or sampling points). A key step is the identification of signals from chemically unknown molecules (peaks) using specialised informatics tools. After standard data processing, some compounds can be identified, but many remain unknown because their chemical structures are not yet recorded in the database (Blaženović et al., 2019). In an average metabolomic study, not every detected signal can be reliably assigned to a specific metabolite, owing to computational artefacts and the presence of chemical impurities. Moreover, not every unidentified compound is necessarily biologically significant, even if it shows a statistically significant difference. To improve reliability, a concerted effort is required to expand metabolomic databases. For instance, according to the Human Metabolome Data Base (HMDB), more than 4,000 different compounds have been identified in human urine (Millers et al., 2019).
In a targeted approach, the concentrations of a predefined group of known metabolites, or specific metabolites of interest, are measured based on particular research hypotheses (Bingol, 2018). This approach is typically used to quantify known metabolites (e.g., molecular markers of disease, or degradation products of a drug’s active substance) for which there are reference standards and validated protocols. Accurate quantification requires the use of appropriate calibration standards and validated methods. Highly sensitive techniques enable the reliable measurement of very small amounts of metabolites, whereas less sensitive methods may fail to quantify low concentrations accurately. It is important to note that accurate and reliable quantitative data cannot be obtained when metabolite concentrations fall below the method’s limit of quantification (LOQ) (Beger et al., 2024). Consequently, concentrations cannot always be determined for certain identified metabolites. Compared to untargeted profiling, targeted approaches yield simpler metabolomic approaches with higher precision and accuracy. Untargeted approaches are mainly applied to investigate pathophysiological mechanisms, whereas targeted approaches are used to refine diagnoses and validate molecular disease markers for potential clinical implementation (Hajnajafi and Iqbal, 2025). The parameters of LOD and LOQ play a central role in metabolomics, as they determine which metabolites can be detected and quantified in a sample, directly influencing the accuracy and reliability of results. Distinguishing between these two parameters is essential both for selecting suitable analytical methods and for correctly interpreting data, since low metabolite concentrations may not be accurately quantified, or even detectable.
Sample types in metabolomic research
The samples most commonly used in metabolomics research are plasma, serum, and urine, although whole blood, cerebrospinal fluid, saliva, bile, synovial fluid, seminal fluid and faeces can also be analysed. The first metabolomic analysis of the spatial distribution of metabolites across a histological preparation of a mouse kidney tissue section was performed using MALDI-TOF-MS technology (Xu et al., 2023).
Different samples require specific preparation methods to ensure reproducibility and reliability of results. Serum and plasma are among the most commonly used samples in metabolomics research as they are stable, show good repeatability and contain a broad spectrum of metabolites (Jobard et al., 2016). Since they circulate throughout the body, they reflect systemic metabolic changes and provide comprehensive insight into physiological and pathological processes. In addition to allowing standardised sample preparation, they are also readily available, making them highly suitable for clinical and biomedical research. The main difference between serum and plasma lies in the presence of coagulation proteins (e.g., fibrinogen, prothrombin), which are present in the plasma but removed from serum. During blood coagulation eicosanoid synthesis is stimulated, meaning that the actual concentrations of these metabolites cannot be measured in serum as they can in plasma (Yamaguchi et al., 2022). This distinction may influence the interpretation of results. Serum and plasma analyses are particularly useful for biomarker discovery, as they provide a systems-level perspective on metabolic changes associated with diseases such as diabetes, cardiovascular, and neurodegenerative diseases.
Urine is an excellent sample for metabolomic research because it contains fewer lipids and protein complexes than other body fluids (Kalantari and Nafar, 2019), thereby reducing the likelihood of interference during analysis. Urine samples can also be stored for long periods without significant degradation, making them ideal for longitudinal studies (Balhara et al., 2023). One challenge in urine metabolomics, however, is the extremely wide dilution range, which can fall below the LOD of analytical methods. Urine is particularly well suited for identifying biomarkers of kidney diseases, as it directly reflects the microenvironment of the kidney.
In addition to blood and urine, stool and saliva are also important samples in metabolic research (Karu et al., 2018; Gardner et al., 2020). Stool is commonly analysed to study the gut microbiome and its metabolome, enabling investigation of how gut bacteria influence host metabolism. Saliva, on the other hand, offers a non-invasive option for detecting biomarkers of both oral and systemic diseases.
Finally, extracellular vesicles (EVs), such as exosomes and microvesicles, are increasingly recognised as valuable samples in metabolomics because they carry metabolites that reflect the physiological and pathological state of their parent cells (Wu et al., 2022). Metabolomic profiling of EVs provides insight into intercellular communication and offers potential for the discovery of novel disease biomarkers.
Challenges in metabolomic research
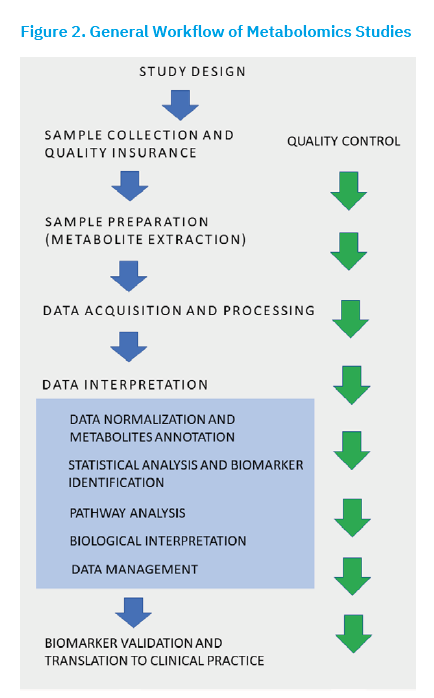
Due to the complexity of the metabolome, which comprises hundreds to thousands of different molecules, even small variations in sample collection or metabolomics workflows (Figure 2) can lead to different results. Each analytical method may produce different outcomes for the same samples, and even instruments with the same configuration from different manufacturers can vary in measurement accuracy and sensitivity, ultimately affecting metabolite detection and quantification. Similarly, the use of different software tools for data processing can lead to variations in results and interpretations. For the further development of metabolomics and its routine application in biomedical research and clinical practise, it is essential to address the major challenges of achieving reproducible, comparable results and standardised procedures (Gonzalez-Dominguez et al., 2020; Thachil et al., 2024). To ensure reproducibility, clear protocols and standards for metabolite analysis must be established, including guidelines for sample preparation, storage and analysis. Factors such as temperature, storage duration, and even the type of reagents or laboratory consumables used can significantly influence outcomes, underscoring the need for strict standardisation. Standardised protocols not only improve consistency but also help minimise errors in metabolomic analysis.
Due to high biological variability between individuals, metabolomics study results can also differ depending on the population being studied, further complicating comparisons across studies. When interpreting results, it is crucial to consider all relevant factors including age, gender, diet, and other lifestyle habits (Navarro et al., 2023). Identifying unknown metabolites and interpreting the biological significance of observed changes is also challenging, particularly given the complexity and variability of metabolomic data. Comparing results among laboratories is difficult due to the lack of standardisation. Variations in techniques and methods for processing raw data, e.g., statistical normalisation methods, as well as the false discovery rate, can affect metabolite detection and analytical outcomes. Ultimately, standardising procedures facilitates comparisons across laboratories, enhances the credibility and reliability of results, and supports the application of metabolomic data in biomedical research and clinical practise.
Sepsis associated acute kidney injury
Sepsis-associated acute kidney injury (SA-AKI) is a heterogeneous syndrome resulting from mechanisms directly related to infection or the host’s response to infection, as well as mechanisms indirectly associated with the consequences of sepsis or its therapies (Zarbock et al., 2023), such as haemodynamic changes, hypoxia, and therapeutic interventions. SA-AKI refers to the occurrence of acute kidney injury (AKI) confirmed according to clinical, biochemical and functional criteria, in the context of sepsis. In human medicine, the KDIGO (Kidney Disease: Improving Global Outcomes) classification is most commonly applied, which defines AKI based on changes in urine output and serum/plasma creatinine concentration. The most common risk factors for SA-AKI include Gram-negative bacteraemia and septic shock, although Gram-positive infections, the use of vasopressors, inhibitors of the renin-angiotensin-aldosterone system, and mechanical ventilation, as well as comorbidities such as chronic liver or kidney disease, hypertension and diabetes, can also contribute to kidney injury. At present, there is no universally accepted definition of SA-AKI in either human or veterinary medicine.
Definitions of SA-AKI vary across studies, and epidemiological data differ depending on diagnostic criteria, study population, and treatment context. A proposed sub-phenotype of SA-AKI is sepsis-induced acute kidney injury (SI-AKI), in which kidney injury results directly from sepsis-related mechanisms. However, this classification is not yet widely accepted in the literature. SI-AKI excludes kidney injury that arises as an indirect consequence of sepsis or its therapy, such as AKI caused by nephrotoxic drugs. Cellular injury mechanisms, including microcirculatory failure and mitochondrial dysfunction, are thought to contribute to the development of SI-AKI (Zarbock et al., 2023). Dysregulation of the host immune response is also an important factor in its pathogenesis.
According to the KDIGO guidelines and Sepsis 3 criteria, SA-AKI in human medicine is defined as AKI that develops within 7 days of the sepsis diagnosis. Numerous studies have shown that SA-AKI carries a worse prognosis than sepsis or AKI alone (Hoste et al., 2015; Peerapornratana et al., 2019). This is attributed to systemic inflammatory changes and more severe organ dysfunction, which leads to a higher incidence of long-term complications. The development of AKI later in the course of sepsis is associated with poorer clinical outcome and higher mortality. To address the issue, the Acute Disease Quality Initiative (ADQI) working group reached a consensus in 2023, proposing that AKI diagnosed within 48 hours of sepsis onset be classified as early SA-AKI, and AKI diagnosed between 48 hours and 7 days as late SA-AKI (Zarbock et al., 2023). However, because sepsis follow-up begins at admission, distinguishing between early and late SA-AKI can be challenging when diagnosis is delayed. Moreover, additional causative factors beyond sepsis itself may contribute to the development of late SA-AKI.
In veterinary medicine, AKI in stages I and II are defined by an increase in serum creatinine concentration of ≥26.4 mmol/L within 48 hours or urine output of less than 1 mL/kg/hour within six hours, according to the criteria of the International Renal Interest Society (IRIS) (Hokamp and Nabity, 2016). The pathophysiological mechanisms of SA-AKI include vascular, immunological and metabolic changes induced by sepsis, which also reduce renal solute clearance. As a result, fluid, toxic metabolites and renally excreted drugs accumulate, accompanied by disturbances in the acid-base balance. The frequency and severity of clearance disorders depend on the degree of AKI and the overall condition of the animal.
Pathophysiology of SA-AKI
Although sepsis requires activation of the host immune system, uncontrolled and generalised inflammation can directly damage organs, leading to the development of acute kidney injury. The pathophysiology of SA-AKI involves excessive inflammation that causes vasodilation, microvascular thrombosis, microcirculatory dysfunction, reduced renal blood flow, oxidative and nitrosative stress, and mitochondrial dysfunction. Together, these processes result in tubular injury and impaired renal function. Cytokines in the bloodstream stimulate endothelial cells to produce nitric oxide (NO), which induces vasodilation and disrupts autoregulation of blood pressure, thereby reducing renal perfusion. Activated macrophages and neutrophils release reactive oxygen species (ROS) and reactive nitrogen species (RNS), further contributing to oxidative and nitrosative stress. In addition, activated endothelium can promote the development of microvascular thrombosis and peripheral haemodynamic disturbances, which aggravate renal hypoperfusion. Disruption of endothelial cell junctions allows intravascular fluid to leak into the intercellular space, resulting in relative hypovolaemia. Reduced renal blood flow (RBF) impairs the delivery of oxygen and metabolic substrates while limiting the removal of cellular by-products such as lactate. According to the latest research findings, the key pathological mechanism of SA-AKI is microcirculation dysfunction, which may occur even in the presence of seemingly adequate renal perfusion and oxygenation (Post et al., 2017).
Renal tubular epithelial cells (RTEC) are responsible for tubular reabsorption and secretion and produce large amounts of ATP. As highly metabolically active cells, RTECs are very sensitive to injury caused by hypoperfusion and dysoxia. Similar to leukocytes, RTECs express Toll-like Receptors (TLR), which recognise pathogen-associated molecular patterns (PAMP) and damage associated molecular patterns (DAMP). Because TLRs are present on both the apical and basolateral membranes, PAMPs and DAMPs can bind after circulating through the peritubular capillaries or filtering through the glomerulus, leading to a double injury of RTECs (Manrique-Caballero et al., 2022). Under these conditions, RTECs shift their priorities, temporarily neglecting specialised resorptive and secretory functions. They switch from oxidative phosphorylation to aerobic glycolysis, using carbohydrates instead of fatty acids as energy substrates. Additional energy is obtained through autophagy, by which the cell degrades damaged organelles, aggregated proteins and pathogens, while recycling metabolic precursors such as nucleotides and amino acids (He et al., 2022). Although this adaptation supports cell survival, it causes additional stress and reduces RTEC functionality.
Sepsis also induces structural and functional changes in kidney cell mitochondria. Reduced oxygen supply diminishes mitochondrial ATP production, while impaired electron transport chain activity promotes oxidative stress and excessive ROS production. Increased mitochondrial membrane permeability leads to swelling, apoptosis and further deterioration of kidney function.
Metabolomic studies of SA-AKI
Although the pathophysiological processes of SA-AKI are being intensively studied, significant gaps remain in the understanding of these mechanisms, which could be addressed using “-omics” approaches. Metabolomic studies provide a novel perspective for analysing metabolic changes that occur in response to various pathological processes, including sepsis. A key factor for successful research is the choice of experimental model, each with specific advantages and limitations. In vitro models, such as cell lines, help to identify pathological cellular processes, while in vivo models allow monitoring of the organism’s actual response to pathological stimuli, thus providing a more comprehensive view of the pathophysiological changes. However, reproducing experimental conditions that accurately mimic sepsis and induce SA-AKI remains challenging, as the specific circumstances leading to renal injury must be recreated within the complexity of a living organism. Metabolomic approaches can significantly improve the understanding of these processes and facilitate the identification of novel biomarkers and potential therapeutic targets for the prevention and treatment of SA-AKI.
Metabolomic research on the pathophysiology of SA-AKIExperimental models of SA-AKIEndotoxaemia
An endotoxin, or lipopolysaccharide (LPS), is an integral component of the outer membrane of Gram-negative bacteria that can trigger a strong inflammatory response in the host organism. It activates the nuclear factor kappa B (NF-kB) signalling pathway and stimulates macrophages (M1) to produce the pro-inflammatory cytokines interleukin 6 (IL-6) and tumour necrosis factor alpha (TNF-a). LPS can also cause renal dysfunction through the activation of mitogen-activated protein kinase (MAPK), inducible NO synthase and cyclooxygenase-2 (COX-2).
Disulphide bond-A oxidoreductase-like protein (DsbA-L) is also involved in the development of AKI by directly stimulating anion-selective channels in proximal tubule cells. In mice with a deletion of the DsbA-L gene, LPS administration did not lead to increased creatinine levels, and M1 macrophage polarisation was reduced. Significantly altered arginine metabolism was detected using untargeted metabolomics with LC-MS/MS (Cui et al., 2023).
Fibroblast growth factor 19 (FGF 19) is a hormone that regulates the biosynthesis of bile acids in the intestine and influences lipid metabolism. Administration of FGF 19 for 7 days protected the kidneys of mice from LPS-induced damage. Targeted metabolomics using LC-MS revealed a decrease in serum fatty acids and malondialdehyde concentration and an increase in catalase activity. Changes in the concentrations of linoleic and g-linolenic acid were also associated with mitochondrial function, regulation of oxidative stress, and ROS formation (Liu et al., 2022).
Systemic Inflammatory Response Syndrome was investigated using a non-targeted metabolomic approach with GC-MS. Fourteen days of low-dose LPS treatment in rats resulted in alterations in the biosynthesis of aminoacyl-tRNA, arginine, and bile acids, and the metabolism of glyoxylate and dicarboxylate, glutathione, glycerolipids, glycine, serine, and threonine. Significant changes were observed in serum for glycine, 5-oxoproline, alanine, glutamic acid, isoleucine, leucine, methionine, proline, serine, threonine, tryptophan, tyrosine and valine (Geng et al., 2020).
Non-targeted GC-MS metabolomics was used to analyse rat serum 2 and 6 hours after LPS administration, which induced oxidative stress, alterations in the citric acid cycle and fatty acid metabolism. Serum levels of docosahexaenoic acid, arachidonic acid, linoleic acid, oleic acid, palmitic acid, and stearic acid gradually increased, while petroselinic acid was elevated only 2 hours after LPS administration. In the renal cortex, changes were observed in the metabolism of taurine, hypotaurine, and phenylalanine, and in the biosynthesis of pantothenate, coenzyme A, and unsaturated fatty acids. Two hours after LPS administration, increased levels of cysteine and taurine, together with decreased levels of b-alanine, pantothenate and uracil induced reduced antioxidant capacity and disturbances in energy and pyrimidine metabolism during the early phase of sepsis. Changes in phenylalanine metabolism, including increased tyrosine, hippuric acid, and phenylacetylglycine levels, suggested a possible link to the stress response in rats (Ping et al., 2019, 2021).
Targeted and non-targeted LC-MS/MS metabolomic studies in SA-AKI mice revealed that 24 hours after administration of a high dose of LPS, the Warburg effect occurs, characterised by disturbances in the citric acid cycle and central carbohydrate metabolism, including impaired transfer of acetyl groups to mitochondria. Catabolic products of polyamines (putrescine, N-acetyl spermidine, N-acetyl putrescine) accumulated in the kidneys of mice. An integrated metabolomics and proteomics approach further revealed changes in the metabolism of alanine, aspartate, glutamate, arginine, proline, histidine, nucleotides, nicotinate, nicotinamide, butanoate, glyoxylate, and dicarboxylate, and also in the biosynthesis of pantothenate and coenzyme A (Xu et al., 2023).
The effect of LPS on the kidneys, lungs, liver and colon of rats was investigated using untargeted metabolomic analysis with UPLC-MS. LPS induced changes in the metabolism of D-glutamine and D-glutamate, taurine and hypotaurine, b-alanine, glutathione, butanoate, and arginine, which are closely linked to the inflammatory response and oxidative stress. The changes affected the metabolism of alanine, aspartate and glutamate, arginine and proline, and cysteine and methionine. Significant alterations in fatty acid metabolism included oleic acid, linoleic acid and arachidonic acid and their metabolites. LPS also caused an increase in cystine and malondialdehyde and a decrease in cysteine, glutamate, aspartate, proline, alanine, tyrosine and isoleucine as well as the antioxidants serine and glycine. Reduced levels of branched-chain amino acids indicate disruptions in energy production and protein synthesis (Gao et al., 2021).
Bacteraemia
Five hours after intravenous administration of Escherichia coli, non-targeted 1H NMR metabolomic analysis revealed decreased levels of lactate, nicotinuric acid, valine, aspartate, glucose, and threonine in the renal tissue of pigs. At the same time, serum valine, glucose and betaine levels decreased, while glutamine levels increased. Elevated lactate, pyruvate and alanine levels indicated reduced involvement of acetyl-coenzyme A in the citric acid cycle. In urine, levels of isovaleroglycine, aminoadipic acid, N-acetyl-glutamine, N-acetyl-aspartate and ascorbate increased, whereas myoinositol and phenylacetyl-glycine decreased. These metabolic alterations included fatty acid metabolism (nicotinuric acid, phenylacetyl-glycine, isovaleroglycine) and b-oxidation of fatty acids in the mitochondria (increased acyl-glycine). Furthermore, serum glucose and betaine, renal tissue valine and aspartate, and urinary phenylacetyl-glycine were significantly correlated with serum creatinine and neutrophil gelatinase-associated lipocalin (Izquierdo-Garcia et al., 2019).
Polymicrobial sepsis
One of the experimental models used in polymicrobial sepsis research is cecal ligation and puncture (CLP), which can induce SA-AKI. The consequences of CLP often include reduced creatinine clearance and decreased antioxidant activity in the kidneys, reflected by increased oxidative stress and elevated levels of TNF-α, IL-1β and TGF-β (transforming growth factor beta).
In polymicrobial sepsis, renal damage in mice was observed as early as 8 hours after CLP, with increased serum levels of cystatin C, urea and creatinine. Untargeted LC-MS/MS metabolomic profiling of renal tissue revealed increased levels of glucose, glycolytic intermediates (2- and 3-phosphoglycerate, pyruvate) and sorbitol, indicating activation of sorbitol metabolism. Decreased levels of citric acid cycle intermediates indicate impaired mitochondrial function, while decreased levels of antioxidants such as ascorbate, α-tocopherol and ergothioneine point to oxidative stress. In response to sepsis, mice also attempted to retain fluid and concentrate urine, as evidenced by reduced betaine levels and increased concentrations of osmolyte (pinitol, urea, taurine) (Waltz et al., 2016).
In addition to renal dysfunction, systemic inflammation is activated 24 hours after CLP in mice. Non-targeted NMR metabolomics identified the involvement of mitochondrial metabolic pathways related to BCAA metabolism, fatty acid oxidation, and de novo nicotinamide adenine dinucleotide (NAD) biosynthesis. Fatty acid oxidation in the renal cortex was inversely correlated with serum creatinine levels. In mouse serum, lactate levels were reduced, while allantoic acid, arabinose and 2,3-butanediol levels were increased, with the latter showing the strongest ability to discriminate between groups (highest VIP value). In urine, levels of ascorbate and metabolites related to BCAA metabolism, ketogenesis and coenzyme A were decreased, while levels of quinolinic acid, glucose, a-ketoisovaleric acid, and 3-methyl-2-oxovaleric acid were increased. These changes affected glycerophospholipid metabolism, glycine, serine and threonine metabolism and ketone metabolism. The metabolites that contributed most to the characterisation of polymicrobial sepsis were glycerophosphocholine, choline, acetoacetic acid, and lactate (Standage et al., 2021).
In a mouse model of polymicrobial sepsis, targeted LC-MS metabolomics was used to investigate the relationship between reduced fatty acid oxidation, proximal tubule cell damage and increased inflammation in mice with peroxisome proliferator-activated receptor alpha (PPARa) deficiency. Significant changes were found in the metabolism of tryptophan-kynurenine-NAD+ and pantothenate in serum. The authors further pointed out that children with sepsis who exhibit suppression of the PPARa signalling pathway often develop a more severe form of AKI (Iwaki et al., 2019).
In sepsis, the permeability of the intestinal barrier is increased, leading to elevated concentration of 4-hydroxyphenyl acetic acid in the serum of patients. Intestinal 4-hydroxyphenylacetic acid can protect the kidneys from SA-AKI by activating the apoptosis repressor with caspase recruitment domain (ARC), which inhibits necroptosis in RTEC. In a targeted LC-MS/MS metabolomic study, treatment with 4-hydroxyphenylacetic acid was found to alleviate renal injury, prevent apoptosis, and increase survival in mice subjected to CLP (An et al., 2024).
Controlled exposure to endotoxin prior to the onset of sepsis, known as preconditioning, can reduce bacterial load in organs and improve survival. Metabolomic analyses using GC-MS and UHPLC-MS/MS of mouse kidney tissue extracts demonstrated that endotoxin preconditioning protects renal macrophages and tubules by reprogramming the citric acid cycle. In unconditioned kidneys exposed to an endotoxin, redox balance was disturbed, meaning that levels of oxidised metabolites increased, while the content of free radicals and compounds that remove them, such as ergothioneine, decreased. At the same time, the content of spermidine and the accumulation of putrescine precursors decreased. By contrast, levels of the anti-inflammatory, antimicrobial itaconate from arginine metabolism increased in the preconditioned kidneys, possibly increasing RTEC resistance. M2 macrophages clustered around the S1 segments of the proximal tubules, but full renal protection required the action of both macrophages and RTECs. Integrated transcriptomic, proteomic and metabolomic analyses revealed that preconditioning in a mouse model of polymicrobial sepsis prevents severe AKI by activating protective stress responses and metabolic pathways in both macrophages and RTECs (Hato et al., 2018).
Spontaneously developed SA-AKI
Non-targeted 1H NMR metabolomic analysis was used to compare children with sepsis and mice with polymicrobial sepsis. In both groups, only a subset developed SA-AKI. According to the research results, the key difference between sepsis and SA-AKI in both children and mice lies in taurine and choline metabolism, specifically in metabolites related to choline metabolism and central carbon metabolism. Since choline administration protected septic mice from developing SA-AKI, the authors suggested that choline treatment may improve renal function in children with sepsis (Hasson et al., 2022).
Untargeted LC-MS/MS metabolomics revealed significant differences in glycerophospholipid, tryptophan, phenylalanine and glycerolipid metabolism between AKI and chronic kidney disease (CKD). In the same study, the first spatial metabolomic analysis of tissue was performed, enabling characterisation of metabolite distribution in the kidneys of mice with AKI. In the renal cortex, amino acids, carnitine, fatty and organic acids were significantly altered, while lipid metabolites showed distinct distributions between the cortex and medulla (Xu et al., 2023). In another GC-MS metabolomic study investigating organ injury in sepsis, the authors concluded that elevated phenylalanine levels could serve as a predictor of AKI risk (Lin et al., 2020).
Using untargeted metabolomic UPLC-MS analysis, differences in the urine metabolome of children with SA-AKI and children with sepsis were assessed at 12 hours and 24 hours. Children with SA-AKI exhibited increased levels of glucose, mannose and lactate, which are sources of glycaemia, and decreased levels of glutamine, a key component of the compensatory citric acid cycle pathway. At 12 hours, significantly altered metabolic pathways included the metabolism of galactose, fructose, mannose, glyoxylate and dicarboxylate, β-alanine, and glycerophospholipid. At 24 hours, tyrosine, ascorbate, and aldarate metabolism were significantly affected. Histidine, indole-3-lactate, trimethylamine-N-oxide and caprylic acid correlated significantly with SA-AKI severity at 12 hours, whereas gentisaldehyde, 3-ureidopropionate, N-4-acetyl-cytidine and 3-methoxy-4-hydroxyphenyl-glycolsulfate were associated at 24 hours. According to the authors, the most effective diagnostic marker for AKI in sepsis, which achieves an area under the curve (AUC) >0.9, is a combination of four statistically significant metabolites at each time point (Wang et al., 2020).
Untargeted GC-MS metabolomics of faecal samples was used to investigate the relationship between indoxyl sulphate, p-cresyl sulphate, phenylacetyl glutamine, and the gut microbiota in early renal dysfunction. Changes in the gut flora associated with these metabolites were associated with the development of early kidney disease. The levels of these metabolites, which are products of bacterial protein fermentation, were inversely correlated with renal function (Barrios et al., 2015).
Metabolomic studies on the effects of drugs on SA-AKI
The effects of baicalin, an extract from the Baikal mussel plant (Scutellariae baicalensis), on LPS-induced AKI were investigated using a 1H NMR metabolomic approach. Mice received baicalin for one week and were sacrificed 24 hours after LPS administration. Analysis of serum and kidney tissue revealed a pronounced inflammatory response, oxidative stress, disruption of energy metabolism, and altered amino acid metabolism. In serum, LPS treatment caused increases in creatinine, NADPH, LDL/VLDL and a decrease in 3-hydroxybutyrate, lactate, alanine, acetate, pyruvate, citrate, taurine, betaine, trimethylamine-N-oxide (TMAO), acetoacetate and glucose. In kidney tissue, LPS induced increases in creatine, creatinine, choline, phosphocholine, TMAO, myoinositol, adenosine and inosine and a decrease in alanine, glutamine, taurine, betaine, lactose, sucrose, maltose, tyrosine, anserine, tryptophan and niacinamide. The administration of baicalin normalised the levels of these metabolites (Liao et al., 2016).
Mice with LPS-induced AKI were treated with the traditional Chinese formulation Huang-Lian-Jie-Du-Decoction (HLJDD). Metabolomic analysis using 1H NMR revealed decreases in choline, phosphocholine and ethanolamine, along with increases in taurine and betaine. HLJDD administration increased glycolysis, i.e., it led to an increase in lactate, alanine and pyruvate in serum and kidney. It also stimulated the citric acid cycle, as indicated by increased citrate in serum and 2-oxoglutarate in the kidneys. At the same time, the levels of ketones (acetoacetate, 3-hydroxybutyrate), creatine and creatinine, as well as the ATP degradation products adenosine and inosine, were decreased. The herbal preparation HLJDD alleviated disruptions in energy metabolism and oxidative stress (Li et al., 2017).
In traditional Chinese medicine, Rehmannia glutinosa Libosch and oleic acid isolated from it are used to treat kidney and inflammatory diseases. A metabolomic approach with NMR showed that oleic acid can alleviate LPS-induced AKI by suppressing inflammation and oxidative stress. Observed metabolic changes included alterations in phenylalanine metabolism (increase in L-tryptophan, L-phenylalanine, phenylalanine), purine metabolism (decrease in L-glutamine, uric acid), sphingolipid metabolism (decrease in phytosphingosine, dihydro-ceramide), and in taurine and hypotaurine metabolism (Zhang et al., 2022).
Egg white protein hydrolysate has a therapeutic effect, including inhibition of pro-inflammatory cytokine production, restoration of redox homeostasis, and alleviation of kidney damage. Metabolomic analysis of rat plasma using UPLC-MS/MS revealed changes in amino acid metabolism, oxidative stress and energy metabolism. In rats treated with LPS, the metabolism of glycine, serine, threonine, arginine, proline, cysteine, methionine, butanoate, phenylalanine, and the citric acid cycle were altered. Egg white protein hydrolysate significantly decreased levels of L-cysteine and BCAA and increased levels of L-cysteine, S-adenosyl-L-methionine (SAMe), glutathione, creatine and citric acid levels (Huan et al., 2022).
Isosteviol-Na, a terpenoid derived from stevioside, exhibits anti-inflammatory, antioxidant and anti-apoptotic effects. Untargeted metabolomic analysis using UHPLC-TIMS-TOF-MS/MS of plasma and kidney tissue revealed significant alterations in glycerophospholipid metabolism. Additionally, arachidonic acid and pantothenate metabolism and coenzyme A biosynthesis were also significantly altered in kidney tissue (Wang et al., 2021).
A metabolomic UHPLC-Q-TOF/MS approach was applied to investigate the influence of the root extract of the coptis plant (Rhizomae coptidis) on mice with SA-AKI. CLP impaired fatty acid oxidation and tryptophan metabolism, and decreased the level of L-carnitine and its metabolites, which normalised after administration of the extract. The extract also reduced ROS production, attenuated renal damage and improved tissue function in mice (Zheng et al., 2021).
Metabolomic 1H NMR analysis showed an increase in creatinine, allantoin and dimethyl-glycine in the urine of SA-AKI rats after CLP. Phenols from ginger, 6- and 10-gingerol decreased 2-oxoglutarate, acetate, allantoin and creatine in urine and increased taurine and antioxidant and anti-inflammatory metabolites: methylsulfonylmethane and dimethylamine. Dimethylglycine and citrate were also significantly altered. Sepsis induced oxidative and nitrosative stress in rat kidneys, whereas gingerol compounds from ginger alleviated renal dysfunction and oxidative/nitrosative stress, reduced pro-inflammatory cytokine levels and increased overall survival (Rodrigues et al., 2018).
Conclusion
Although the designs of metabolomic studies on SA-AKI differ, their conclusions regarding the pathophysiology of SA-AKI largely align with findings from conventional studies. The most commonly observed changes include increased inflammatory signalling, intracellular metabolic changes, oxidative stress and osmolyte imbalances. While individual metabolites may serve as diagnostic markers, they cannot fully capture the complex metabolic changes occurring in patients with SA-AKI.
References [… show]
Metabolomika u istraživanjima akutne ozljede bubrega povezane sa sepsom
Dora Ivšić Škoda1, divsicskoda@vef.unizg.hr; Dina Rešetar Maslov1* (dopisni autor), drmaslov@vef.unizg.hr, orcid.org/0000-0002-0159-8281; Josipa Kuleš2, jkules@vef.unizg.hr, orcid.org/0000-0002-4945-4690; Valerija Benko3, vbenko@vef.unizg.hr, orcid.org/0000-0002-3307-9582; Vlasta Đurić1 vduric@vef.unizg.hr; Renata Barić Rafaj2, rrafaj@vef.unizg.hr, orcid.org/0000-0002-1043-1176; Vladimir Mrljak1, vmrljak@vef.unizg.hr, orcid.org/0000-0002-0723-7815.
1 Klinika za unutarnje bolesti, Veterinarski fakultet Sveučilišta u Zagrebu, 10000 Zagreb, Hrvatska
2Zavod za kemiju i biokemiju, Veterinarski fakultet Sveučilišta u Zagrebu, 10000 Zagreb, Hrvatska
3Zavod za biologiju i patologiju riba i pčela, Veterinarski fakultet Sveučilišta u Zagrebu, 10000 Zagreb, Hrvatska
Metabolomika je znanstvena disciplina usmjerena na analizu malih molekula, takozvanih metabolita, u biološkim uzorcima s ciljem razumijevanja fizioloških i patoloških procesa. Razvoj analitičkih tehnologija poput spektrometrije masa i nuklearne magnetske rezonancije omogućio je napredak u neciljanoj i ciljanoj metabolomskoj analizi. U neciljanoj analizi proučava se širok spektar metabolita bez prethodnih pretpostavki, dok se ciljana analiza usredotočuje na kvantifikaciju poznatih molekula. Biološki uzorci poput seruma, plazme i urina najčešće se koriste, a izbor ovisi o kliničkom kontekstu i prirodi bolesti. Unatoč svom potencijalu, metabolomika se suočava s izazovima poput tehničke složenosti, biološke varijabilnosti, nedostatka standardizacije te potrebe za naprednim statističkim metodama. Međutim, u posljednjem desetljeću našla je primjenu u proučavanju sepse i povezane akutne ozljede bubrega, stanja s visokom smrtnošću i ograničenim terapijskim mogućnostima. U ovom preglednom radu prikazano je što metabolomika, s kojim se izazovima suočava i trenutačno stanje istraživanja metabolomike u kontekstu sepsom povezane akutne ozljede bubrega (SA-AKI). Metabolomička istraživanja SA-AKI identificirala su potencijalne ključne biološke promjene poput: pojačane upalne signalizacije, oksidativnog stresa, poremećaja osmolita i staničnih metaboličkih disfunkcija. Iako su pojedinačni metaboliti prepoznati kao potencijalni dijagnostički markeri, niti jedan ne može u potpunosti obuhvatiti složene metaboličke promjene karakteristične za SA-AKI. U budućnosti bi integracija metabolomike s drugim „-omikama“ i naprednim analitičkim pristupima mogla unaprijediti rano otkrivanje bolesti, razumijevanje patofiziologije i razvoj personaliziranih terapija.
Ključne riječi: metabolomika, metabolom, metaboliti, akutna ozljeda bubrega, sepsa

